 |
 |
| Anim Biosci > Volume 32(10); 2019 > Article |
|
Abstract
Objective
Methods
Results
Conclusion
ACKNOWLEDGMENTS
Figure 1
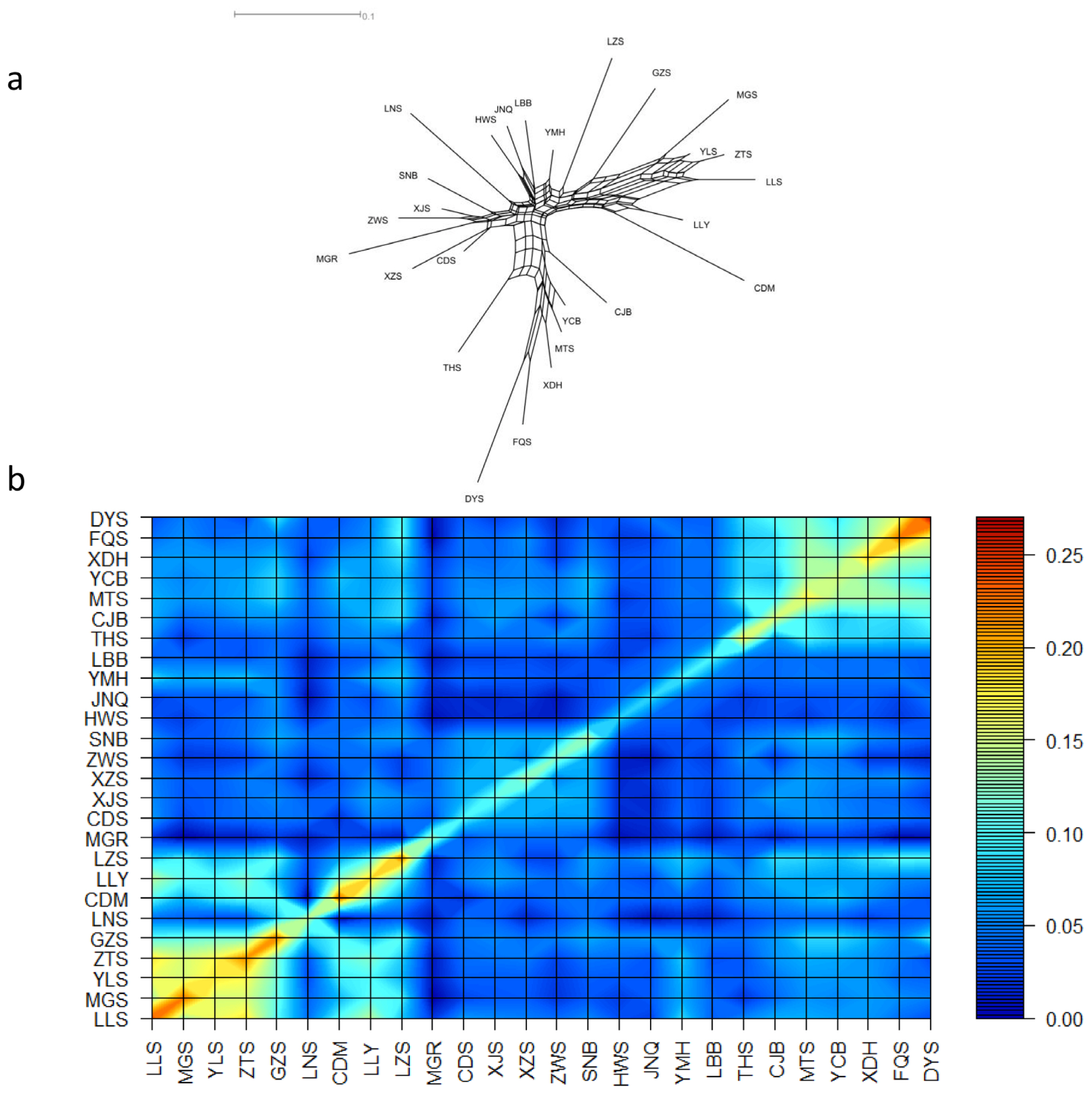
Table 1
| Population1) | MNA±SD | NEA±SD | HE±SD | HO±SD | Rt±SD | FIS | NE(0.05) |
|---|---|---|---|---|---|---|---|
| LLS | 6.00±2.56 | 2.932±0.315 | 0.588±0.030 | 0.491±0.013 | 5.22±2.13 | 0.167 | 120.8 |
| MGS | 5.33±1.95 | 2.847±0.189 | 0.598±0.033 | 0.573±0.013 | 4.7±1.56 | 0.043 | 74.1 |
| YLS | 6.00±2.68 | 3.162±0.261 | 0.628±0.031 | 0.552±0.013 | 5.36±2.16 | 0.123 | 171.3 |
| ZTS | 5.93±2.52 | 2.88±0.225 | 0.593±0.034 | 0.530±0.013 | 5.17±1.98 | 0.107 | 97.1 |
| GZS | 5.63±2.27 | 2.77±0.227 | 0.582±0.032 | 0.546±0.013 | 4.91±1.88 | 0.062 | 113.6 |
| LNS | 6.47±2.74 | 3.528±0.245 | 0.664±0.034 | 0.619±0.012 | 5.72±2.26 | 0.066 | 153.7 |
| CDM | 5.93±2.41 | 2.877±0.2 | 0.606±0.028 | 0.595±0.012 | 5.11±1.79 | 0.018 | 80.4 |
| LLY | 5.63±2.22 | 3.131±0.216 | 0.64±0.026 | 0.600±0.013 | 5.13±1.82 | 0.062 | 74.4 |
| LZS | 5.47±2.26 | 2.793±0.219 | 0.583±0.034 | 0.539±0.012 | 4.71±1.77 | 0.075 | 75.9 |
| MGR | 6.93±2.63 | 3.664±0.293 | 0.661±0.037 | 0.556±0.013 | 6.19±2.3 | 0.161 | 143.1 |
| CDS | 7.90±3.08 | 4.102±0.291 | 0.709±0.029 | 0.653±0.011 | 6.87±2.47 | 0.080 | 124.1 |
| XJS | 8.23±2.93 | 3.962±0.29 | 0.689±0.036 | 0.616±0.012 | 7.03±2.27 | 0.108 | 101.2 |
| XZS | 7.37±3.10 | 3.625±0.239 | 0.685±0.027 | 0.612±0.012 | 6.61±2.49 | 0.108 | 96.1 |
| ZWS | 6.87±2.69 | 3.903±0.303 | 0.694±0.035 | 0.622±0.018 | 6.8±2.66 | 0.106 | 569.7 |
| SNB | 6.50±2.33 | 3.685±0.264 | 0.684±0.028 | 0.600±0.012 | 5.86±1.98 | 0.121 | 47.5 |
| HWS | 7.37±2.28 | 4.294±0.312 | 0.729±0.025 | 0.588±0.013 | 6.75±1.96 | 0.195 | 113.2 |
| JNQ | 7.30±2.51 | 4.369±0.299 | 0.737±0.024 | 0.646±0.012 | 6.56±2.12 | 0.124 | 98.3 |
| YMH | 7.83±2.77 | 3.859±0.299 | 0.704±0.023 | 0.557±0.013 | 6.73±2.24 | 0.210 | 66.1 |
| LBB | 7.67±2.51 | 3.822±0.234 | 0.707±0.025 | 0.628±0.013 | 6.72±2.07 | 0.113 | 76.7 |
| THS | 6.47±2.34 | 3.344±0.237 | 0.657±0.027 | 0.615±0.011 | 5.59±1.95 | 0.022 | 134.7 |
| CJB | 5.77±2.22 | 3.275±0.26 | 0.644±0.029 | 0.623±0.014 | 5.32±1.97 | 0.032 | 374.2 |
| MTS | 5.37±2.09 | 3.014±0.224 | 0.625±0.025 | 0.569±0.012 | 4.78±1.82 | 0.091 | 69.4 |
| YCB | 5.93±2.32 | 3.25±0.23 | 0.655±0.024 | 0.623±0.012 | 5.19±1.88 | 0.050 | 100.1 |
| XDH | 5.37±2.24 | 2.912±0.225 | 0.608±0.030 | 0.549±0.013 | 4.74±1.81 | 0.097 | 96.8 |
| FQS | 4.90±2.40 | 2.723±0.247 | 0.569±0.034 | 0.475±0.012 | 4.30±1.93 | 0.168 | 89 |
| DYS | 4.40±2.25 | 2.424±0.217 | 0.507±0.039 | 0.434±0.013 | 3.89±1.91 | 0.146 | 32.7 |
| Average | 6.33±2.47 | - | 0.644±0.030 | 0.577±0.013 | 5.61±2.04 | 0.167 | - |
n, sample size; MNA, mean number of alleles; SD, standard deviation; NEA, mean number of effective alleles; HE, expected heterozygosity; HO, observed heterozygosity; Rt, allelic richness; FIS, fixation index; NE(0.05), effective population size based on linkage disequilibrium (minor allele frequency 0.05).
1) LLS, Longling yellow goat; MGS, Maguan poll goat; YLS, Yuling goat; ZTS, Zhaotong goat; GZS, Guizhou White goat; LNS, Liaoning Cashmere goat; CDM, Chengdu Brown goat; LLY, Longlin goat; LZS, Leizhou goat; MGR, Inner Mongolia Cashmere goat; CDS, Chaidamu goat; XJS, Xinjiang goat; XZS, Tibetan goat; ZWS, Zhongwei goat; SNB, Shannan White goat; HWS, Huanghuai goat; JNQ, Jining Gray goat; YMH, Yimeng Black goat; LBB, Lubei White goat; THS, Taihang goat; CJB, Yangtse River Delta White goat; MTS, Matou goat; YCB, Yichang White goat; XDH, Xiangdong Black goat; FQS, Fuqing goat; DYS, Daiyun goat.
Table 2
| Breeds1) | WEDs | Bootstrap | WLM | PCHe | PCWeitz | PCFst | PC5:1 |
|---|---|---|---|---|---|---|---|
| LLS | 0 | 0 | 0 | −0.047 | 2.81 | 0.349 | 2.333 |
| MGS | 0.0243 | 0.0211 | 0 | −0.007 | 3.76 | 0.515 | 3.131 |
| YLS | 0.0073 | 0.0143 | 0.0419 | −0.035 | 1.58 | 0.189 | 1.310 |
| ZTS | 0 | 0 | 0 | −0.098 | 1.96 | 0.187 | 1.616 |
| GZS | 0 | 0 | 0 | −0.161 | 2.67 | 0.232 | 2.197 |
| LNS | 0.1361 | 0.1324 | 0.1264 | 0.258 | 3.71 | 0.736 | 3.133 |
| CDM | 0.0048 | 0.0063 | 0.0242 | 0.121 | 6.71 | 1.034 | 5.610 |
| LLY | 0 | 0 | 0 | −0.047 | 3.71 | 0.473 | 3.083 |
| LZS | 0 | 0 | 0 | −0.148 | 6.57 | 0.783 | 5.448 |
| MGR | 0.1622 | 0.1613 | 0.1071 | 0.307 | 7.00 | 1.235 | 5.882 |
| CDS | 0.0229 | 0.0227 | 0.0964 | 0.173 | 2.81 | 0.538 | 2.370 |
| XJS | 0 | 0 | 0 | 0.071 | 2.86 | 0.457 | 2.394 |
| XZS | 0.0352 | 0.0255 | 0.0406 | 0.227 | 5.45 | 0.951 | 4.578 |
| ZWS | 0.0877 | 0.0972 | 0.0781 | 0.074 | 2.79 | 0.450 | 2.336 |
| SNB | 0 | 0 | 0 | 0.086 | 3.66 | 0.581 | 3.063 |
| HWS | 0.2017 | 0.2027 | 0.0853 | 0.223 | 5.6 | 0.969 | 4.702 |
| JNQ | 0.1488 | 0.1469 | 0.2621 | 0.363 | 2.21 | 0.619 | 1.902 |
| YMH | 0 | 0 | 0 | 0.035 | 2.04 | 0.313 | 1.705 |
| LBB | 0.1353 | 0.1296 | 0.1007 | 0.182 | 4.56 | 0.789 | 3.829 |
| THS | 0 | 0 | 0.0181 | −0.014 | 1.94 | 0.256 | 1.614 |
| CJB | 0 | 0 | 0 | −0.071 | 3.34 | 0.402 | 2.770 |
| MTS | 0 | 0 | 0 | −0.195 | 3.49 | 0.316 | 2.875 |
| YCB | 0 | 0 | 0 | −0.124 | 4.14 | 0.467 | 3.428 |
| XDH | 0 | 0 | 0 | −0.218 | 2.52 | 0.161 | 2.063 |
| FQS | 0.0337 | 0.04 | 0.0193 | −0.200 | 2.67 | 0.198 | 2.191 |
| DYS | 0 | 0 | 0 | −0.207 | 3.49 | 0.305 | 2.873 |
Contribution made by each breed to total genetic diversity for 26 Chinese indigenous goat breeds based on methods.
MEK, marker-estimated kinships; WEDs, which vary based on weighted equal drift similarity; Bootstrap, WEDS with bootstrap procedure; WLM, weighted log-linear model; PCweitz, Weitzman approach; PCHe, proportion of expected heterozygosity; PCFst, aggregate methods based on Fst; and PC5:1, the Piyasation and Kinghorn formula. Values representing high contributions to genetic diversity are shown in boldface.
1) LLS, Longling yellow goat; MGS, Maguan poll goat; YLS, Yuling goat; ZTS, Zhaotong goat; GZS, Guizhou White goat; LNS, Liaoning Cashmere goat; CDM, Chengdu Brown goat; LLY, Longlin goat; LZS, Leizhou goat; MGR, Inner Mongolia Cashmere goat; CDS, Chaidamu goat; XJS, Xinjiang goat; XZS, Tibetan goat; ZWS, Zhongwei goat; SNB, Shannan White goat; HWS, Huanghuai goat; JNQ, Jining Gray goat; YMH, Yimeng Black goat; LBB, Lubei White goat; THS, Taihang goat; CJB, Yangtse River Delta White goat; MTS, Matou goat; YCB, Yichang White goat; XDH, Xiangdong Black goat; FQS, Fuqing goat; DYS, Daiyun goat.
Table 3
| Breed2) | f ii | DNei | Contribution to f | Contribution to D | GDT|i | Loss/gain (%) | PC1 (%) | PC2 (%) |
|---|---|---|---|---|---|---|---|---|
| LLS | 0.4165 | 0.1232 | 0.0108 | 0.0272 | 0.7408 | 0 | 3.673 | 3.673 |
| MGS | 0.4078 | 0.1218 | 0.0102 | 0.0264 | 0.7406 | 0 | 3.565 | 3.700 |
| YLS | 0.3781 | 0.1053 | 0.0091 | 0.0253 | 0.7408 | 0 | 3.417 | 3.767 |
| ZTS | 0.4130 | 0.1166 | 0.0105 | 0.0260 | 0.7412 | 0.1 | 3.511 | 3.646 |
| GZS | 0.4236 | 0.1156 | 0.0110 | 0.0256 | 0.7417 | 0.2 | 3.457 | 3.592 |
| LNS | 0.3439 | 0.1123 | 0.0093 | 0.0330 | 0.7386 | −0.3 | 4.456 | 3.997 |
| CDM | 0.4001 | 0.1279 | 0.0111 | 0.0313 | 0.7396 | −0.1 | 4.227 | 3.781 |
| LLY | 0.3678 | 0.0990 | 0.0099 | 0.0282 | 0.7409 | 0 | 3.808 | 3.794 |
| LZS | 0.4233 | 0.1177 | 0.0121 | 0.0288 | 0.7416 | 0.1 | 3.889 | 3.605 |
| MGR | 0.3442 | 0.1222 | 0.0074 | 0.0276 | 0.7382 | −0.3 | 3.727 | 4.037 |
| CDS | 0.2967 | 0.0829 | 0.0086 | 0.0343 | 0.7392 | −0.2 | 4.632 | 4.078 |
| XJS | 0.3159 | 0.0841 | 0.0094 | 0.0335 | 0.7400 | −0.1 | 4.524 | 3.983 |
| XZS | 0.3217 | 0.1014 | 0.0081 | 0.0306 | 0.7388 | −0.2 | 4.132 | 4.051 |
| ZWS | 0.3217 | 0.0961 | 0.0040 | 0.0143 | 0.7400 | −0.1 | 1.931 | 4.010 |
| SNB | 0.3218 | 0.0884 | 0.0090 | 0.0318 | 0.7399 | −0.1 | 4.294 | 3.983 |
| HWS | 0.2780 | 0.0808 | 0.0067 | 0.0297 | 0.7389 | −0.2 | 4.011 | 4.172 |
| JNQ | 0.2692 | 0.0840 | 0.0073 | 0.0356 | 0.7378 | −0.4 | 4.808 | 4.240 |
| YMH | 0.3038 | 0.0752 | 0.0083 | 0.0303 | 0.7403 | 0 | 4.092 | 4.010 |
| LBB | 0.3011 | 0.0889 | 0.0071 | 0.0280 | 0.7392 | −0.2 | 3.781 | 4.091 |
| THS | 0.3486 | 0.0928 | 0.0107 | 0.0331 | 0.7406 | 0 | 4.470 | 3.875 |
| CJB | 0.3641 | 0.0940 | 0.0081 | 0.0227 | 0.7411 | 0.1 | 3.065 | 3.794 |
| MTS | 0.3806 | 0.0946 | 0.0120 | 0.0317 | 0.7419 | 0.2 | 4.281 | 3.713 |
| YCB | 0.3522 | 0.0849 | 0.0098 | 0.0282 | 0.7414 | 0.1 | 3.808 | 3.808 |
| XDH | 0.3976 | 0.0986 | 0.0110 | 0.0270 | 0.7421 | 0.2 | 3.646 | 3.646 |
| FQS | 0.4361 | 0.1197 | 0.0123 | 0.0278 | 0.7420 | 0.2 | 3.754 | 3.551 |
| DYS | 0.4985 | 0.1466 | 0.0119 | 0.0226 | 0.7420 | 0.2 | 3.052 | 3.362 |
f ii, average co-ancestries; DNei, Nei’s genetic distance; f, contribution to global co-ancestry; D, absolute contribution to the total genetic diversity; GDT|i, global diversity; loss/gain(%), the % loss/gain after removing a population from the pool; PC, proportional contribution to gene diversity; PC1 estimates are weighted by population size; PC2 estimates ignore sample size.
1) Values representing high contributions are shown in boldface. Mean co-ancestry within-breed, f = 0.363; mean Nei’s minimum distance in the metapopulation, D = 0.103; mean co-ancestry in the metapopulation, f = 0.246; global genetic diversity of the metapopulation, GDT = 0.741.
2) LLS, Longling yellow goat; MGS, Maguan poll goat; YLS, Yuling goat; ZTS, Zhaotong goat; GZS, Guizhou White goat; LNS, Liaoning Cashmere goat; CDM, Chengdu Brown goat; LLY, Longlin goat; LZS, Leizhou goat; MGR, Inner Mongolia Cashmere goat; CDS, Chaidamu goat; XJS, Xinjiang goat; XZS, Tibetan goat; ZWS, Zhongwei goat; SNB, Shannan White goat; HWS, Huanghuai goat; JNQ, Jining Gray goat; YMH, Yimeng Black goat; LBB, Lubei White goat; THS, Taihang goat; CJB, Yangtse River Delta White goat; MTS, Matou goat; YCB, Yichang White goat; XDH, Xiangdong Black goat; FQS, Fuqing goat; DYS, Daiyun goat.
Table 4
WEDs, which vary based on weighted equal drift similarity; Bootstrap, WEDS with bootstrap procedure; WLM, weighted log-linear model; PCHe, proportion of expected heterozygosity; PCweitz, Weitzman approach; PCFst, aggregate methods based on Fst; and PC5:1, the Piyasation and Kinghorn formula. PC, proportional contribution to gene diversity; PC1 estimates are weighted by population size; PC2 estimates ignore sample size.
REFERENCES
- TOOLS
-
METRICS

- Related articles
-
Evaluation of the genetic diversity of six Chinese indigenous chickens2020 October;33(10)
Diversity of Chinese Indigenous Goat Breeds: A Conservation Perspective - A Review -2004 May;17(5)
Estimates of genetic parameters of some growth traits in jersey cattle1995 December;8(6)






 PDF Links
PDF Links PubReader
PubReader ePub Link
ePub Link Full text via DOI
Full text via DOI Full text via PMC
Full text via PMC Download Citation
Download Citation Supplement
Supplement Print
Print




