 |
 |
Abstract
An in vitro experiment was conducted to evaluate the effect of methylcellulose on the attachment of major cellulolytic bacteria on rice straw and its digestibility. Rice straw was incubated with ruminal mixture with or without 0.1% methylcellulose (MC). The attachment of F. succinogenes, R. flavefaciens and R. albus populations on rice straw was measured using real-time PCR with specific primer sets. Methylcellulose at the level of 0.1% decreased the attachment of all three major cellulolytic bacteria. In particular, MC treatment reduced (p<0.05) attachment of F. succinogenes on rice straw after 10 min of incubation while a significant reduction (p<0.05) in attachment was not observed until 4 h incubation in the case of R. flavefaciens and R. albus. This result indicated F. succinogenes responded to MC more sensitively and earlier than R. flavefaciens and R. albus. Dry matter digestibility of rice straw was subsequently inhibited by 0.1% MC, and there was a significant difference between control and MC treatment (p<0.05). Incubated cultures containing MC had higher pH and lower gas production than controls. Current data clearly indicated that the attachment of F. succinogenes, R. flavefaciens and R. albus on rice straw was inhibited by MC, which apparently reduced rice straw digestion.
Rumen fermentation is unique in that efficient degradation of rough forage such as rice straw relies on the cooperation between microbial consortia and factors in rumen. Under a normal feeding system, bacterial populations associated with feed particles (compartment 2 and 3 microbes in the rumen, as proposed by Czerkawski (1986): bacteria loosely adherent to the solid phase and bacteria firmly adherent to the solid phase, respectively) are numerically predominant and account for 70 to 80% of the total microbial population and microbial ATP production in the rumen (Craig et al., 1987; Minato et al., 1993; Miron et al., 2001). The populations are responsible for most enzyme activities which are 80% of the endoglucanase activity, 70% of the amylase activity and 75% of the protease activity in the rumen (Minato et al., 1966; Brock et al., 1982; McAllister et al., 1994). Fibrolytic activity, hemicellulase and cellulase, is higher in solid-associated populations than that in ruminal fluid (Williams and Strachan, 1984; Michalet-Doreau et al., 2001). These data suggested that microbial populations associated with feed particle digestion played an important role in feed digestion in the rumen. Also, direct microscopic examination clearly showed that forage and grain particles from the rumen are colonized by morphologically diverse microbial populations (Dinsdale et al., 1978; McAllister et al., 1990). The importance of attachment in cellulose digestion and various factors affecting its efficacy were mentioned in previous studies (McAllister et al., 1994; Miron et al., 2001; Marinucci et al., 2006; Sung et al., 2007; Hwang et al., 2008). Mutant cells lacking the adhesion ability had a smoother appearance of surface topology compared to the wild type cells and had lost most of their cellulolytic capability (Gong and Forsberg, 1989; Stewart et al., 1990; Miron and Forsberg, 1998, 1999; Reddy and Morrison, 1998). The previous studies reported that the necessity of attachment for subsequent cellulose degradation by ruminal cellulolytic bacteria was confirmed by using methylcellulose which prevents the attachment of cellulolytic bacteria to cellulose fibers and consequently inhibited cellulose digestion (Minato and Suto, 1979; Pell and Schofield, 1993). Kudo et al. (1987) also reported that treatment of Fibrobacter succinogenes, Ruminococcus flavefaciens, and Ruminococcus albus with 0.1 methylcellulose inhibited filter paper digestion completely, but inhibited endoglucanase activity (IU) by less than that 20%. Major cellulolytic bacteria are well known as firmly attached species (compartment 3 microbes) that can adhere to cellulose during cellulose digestion and their attachments to fibrous particles are inhibited by MC (Methylcellulose: Minato et al., 1993; Pell and Schofield, 1993; McAllister et al., 1994; Miron et al., 2001). However, this is not enough direct evidence on the relationship between MC, bacterial attachment and fiber digestion in the rumen microbial ecosystem, because it was based on microscopic observation, solid and liquid fractionized works and some bacterial pure cultivation. Further studies with molecular techniques were needed to monitor each cellulolytic bacterial strain such as F. succinogenes, R. flavefaciens, and R. albus, during ruminal fermentation.
The objective of this in vitro study was to evaluate the effect of MC on cellulolytic processes and cellulolytic bacterial attachments to rice straw during ruminal fermentation using real time PCR.
Rumen contents were collected from three rumen fistulated Holstein steers (740±10 kg body weight) after morning feeding. The animals were fed twice daily at 09:00 and 17:00 with a mixture of 40% concentrate and 60% timothy hay. Rumen fluid containing the ingesta was subjected to oxygen free CO2 using a gassing apparatus, homogenized with a mixer (Mini mixer, Hanil, Korea) for 1 min, then filtered through 8-layers of cheese cloth to obtain the rumen microbial mixes. Rice straw was grounded to pass a 2 mm screen for this experiment.
Cellulolytic bacterial attachment to rice straw and its digestion in the presence of MC (Methylcellulose) addition was evaluated using an in vitro culture system. The rumen fluids-basal medium (McDougall, 1948) mixture was prepared by mixing one volume of rumen fluid and two volumes of the basal medium. Sixty mL of the rumen fluid-basal medium mixture was inoculated into a 125 mL of serum bottle containing 0.5 g of rice straw under flushing with oxygen free CO2, and then 0.1% (w/v) of MC solution was added to the MC treatment groups. MC solution was prepared using methylcellulose powder (Sigma Mo262) agitated and boiled until the particles were thoroughly wetted and evenly dispersed for complete solubilization. The serum bottles were held in a shaking incubator with 120 rpm at 39°C for 0, 10 min, 2, 4, 8, 12, 24 and 48 h.
Sample preparation: The incubated culture was centrifuged at 160×g for 10 min to separate rice straw and culture medium. Collected rice straw was suspended in 50 mL of 0.9% saline solution and centrifuged three times at 160×g for 10 min to remove detachable bacteria. After centrifugation, rice straw was dried using a lyophilizer (Ilshin, Korea) and kept at −80°C until bacterial measurement.
DNA extraction: gDNA was extracted according to the method described by Purdy et al. (1996). Briefly, 0.5 g of dried rice straw was mixed with 0.35 mL of TE buffer (10 mM Tris HCl, 1 mM EDTA, pH 8.0), 0.5 mL of Tris-buffered phenol and 0.25 g of sterilized glass beads (0.5 mm, BioSpec. Product Inc. USA). The tubes were shaken for 2 min, stood on ice for 2 min and this step was repeated three times. After adding 40 μL of 10% sodium lauryl sulfate solution, tubes were centrifuged at 13,000×g for 2 min and supernatant was collected. The remaining pellet was resuspended in 20 μL of TE buffer, then centrifuged at 13,000×g for 2 min and the supernatant was collected. gDNA was collected from pooled supernatant using a hydroxyapatite chromatography column (Hydroxyapatite Bio-Gel HTP Gel, Bio-Rad Laboratories, Inc, USA). The RNA was removed by DNAse-free pancreatic RNAse Following this treatment there was a subsequent gel filtration (MicroSpin S-200 HR Columns, Amersham Biosciences, UK). The purity and concentration of total DNA were checked using a Biomate 5 spectrophotometer (Thermo Spectronic, USA).
PCR primer: Species-specific PCR primer sets for F. succinogenes, R. flavefaciens and R. albus were selected from the previous study (Koike and Kobayashi, 2001). Primers for F. succinogenes, R. flavefaciens, and R. albus were: Fs219f (5′-GGT ATG GGA TGA GCT TGC-3′) and Fs654r (5′-GCC TGC CCC TGA ACT ATC -3′); Rf154f (5′- TCT GGA AAC GGA TGG TA-3′) and Rf425r (5′-CCT TTA AGA CAG GAG TTT ACA A-3′); Ra1281f (5′-CCC TAA AAG CAG TCT TAG TTC G-3′) and Ra1439r (5′-CCT CCT TGC GGT TAG AAC A-3′), respectively. Amplification sizes from PCR reactions for the three bacterial species were 446, 259 and 175 bp and annealing temperatures were 62, 55 and 55°C, respectively.
Real-time PCR: gDNA was amplified and quantified with an iCycler iQ real-time PCR system (Bio-Rad Inc. USA). The iQ Syber Green Supermix (Bio-Rad INC. USA) was used for real-time PCR (RT-PCR) amplification according to the manufacturer’s protocol. RT-PCR conditions were: one cycle of initial denaturation at 95°C for 3 min, 40 cycles of denaturation at 95°C for 30 s, followed by annealing at each temperature of strains for 30 s and then an extension at 72°C for 30 s. Thereafter, the melting point of RT-PCR product was analyzed to detect specificity of application. The melting curve was obtained by a 0.1°C/s increase of heating temperature from 65 to 95°C with fluorescence detection at 0.1°C intervals. Bacterial population was defined as log copy number of 16S rDNA which was calculated from a standard curve of control plasmid. The control plasmid had an insert of a specific fragment of 16S rDNA amplified with primers specific to each species (F. succinogenes, R. flavefaciens and R. albus). The control plasmid was constructed by using the pGEM-T and pGEM-T Easy Vector System (Promega, USA) according to the manual procedure. The standard curves were respectively made by plotting Ct values for serial dilutions of the each control plasmid for each species.
The DM digestibility was calculated by the difference between dry matter before and after incubation. The change in pH was measured using a Mettler Delta 340 pH meter (Mettler Electronics, UK). The accumulated head gas pressure was measured using a pressure transducer and recorded using a digital readout voltmeter (Laurel Electronics. USA).
All experimental cultures were done in triplicate. Data were analyzed using ANOVA procedure of Statistical Analysis System Institute, Inc. (SAS, 1996). When the overall treatment effect was significant (p<0.05), the differences between treatment means were tested using the least significant difference (LSD) procedure of SAS (1996).
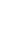
The attachment for subsequent cellulose degradation is an indispensable process in ruminal cellulolytic bacteria but influenced by various factors which is related to bacteria and substrate (Miron et al., 2001). We did focus on the MC related substrate effectors and obtained the clear results of its respective effects on F. succinogenes, R. flavefaciens, and R. albus, as shown in Figure 1, 2 and 3. As in earlier studies (Cheng et al., 1983; Craig et al., 1987) ruminal microorganisms rapidly associated with and attached to ingested feed particles, the attachment of three major cellulolytic bacteria to rice straw took place at the initial time of incubation (0 h) regardless MC addition. However, all of bacterial strains did apparently show the trend of attachment inhibition with different characteristics depending on respective bacterial strains response to the addition of 0.1% MC in accordance to the incubation time. The attachment of major cellulolytic bacteria in the absence of MC gradually increased to reach the maximal level at 24 h incubation and then slowly decreased after that time. This result was similar with the trend seen in normal rumen in vitro culture (Sung et al., 2007). However, as shown in Figure 1, F. succinogenes attachment in the presence of MC was at significantly lower levels compared with control after 10 min of incubation (p<0.05) and thereafter showed a continuous decrease during incubation. In case of R. flavefaciens (Figure 2) and R. albus (Figure 3), their attachment was not different between control and MC addition the earlier stage of incubation (0 to 4 h). However, the attachments of both strains abruptly dropped after 4 h of incubation and stayed significantly lower with MC addition compared with control during the incubation (p<0.05). These results using real-time PCR to monitor attached populations of cellulolytic bacterial suggested that attachment is essential for subsequent fiber digestion in rumen fermentation. Although an earlier study (Williams and Strachan, 1984) had shown a similar result using microscopic observationof solid and liquid fractionized samples from pure bacterial cultivation. Minato and Suto (1978) mentioned that bacteria associated with the solid fraction were important in feed particle digestion. Kudo et al. (1987) using electron microscopy reported that the presence of MC prevents the attachment of cellulolytic rumen bacteria to cellulose fibers. More specifically, R. flavefaciens FD-1 adherence to cellulose, has been shown to be inhibited by MC and carboxylmethyl cellulose (CMC), which suggests that receptors responsible for adherence are blocked by these derivatives (Rasmussen et al., 1989). Also, when attachment on rice straw during early incubation is considered, we find that F. succinogenes was more sensitive to MC treatment than R. flavefaciens and R. albus (Figures 1 and 4). This result was different from the study of Bhat et al. (1990) where the degree of inhibition by MC was similar for the both species, F. succinogenes and R. flavefaciens (about 33%), but inhibition by CMC was greater for F. succinogenes (31%) than R. flavefaciens (24%). As an advance on the earlier studies, our present data obtained with the recent technique of real-time PCR quantification clearly show that attachment to rice straw of major cellulolytic bacteria, F. succinogenes, R. flavefaciens, and R. albus is significantly inhibited by 0.1% MC addition in rumen mixed bacterial digestion. Also, these results showed that inhibition occurred in the three major cellulolytic bacteria during the initial stage of incubation and more sensitively in F. succinogenes than R. flavefaciens and R. albus.
The DM digestibility of rice straw is presented in Figure 5. The DM digestibility was not different between control and MC addition at initial incubation time (0 h), and then became significantly different through the increasing incubation time. The cellulolytic bacterial species such as F. succinogenes, R. flavefaciens, and R. albus play a major role in fiber digestion in the rumen (Lee et al., 2000). Previous studies (Forsberg et al., 2000; Miron et al., 2001) reported that the attachments of bacteria on fiber surface possess cellulolytic enzymes for efficient breakdown of plant cell wall polysaccharide. Similar to our result (Figure 4), Rasmussen et al. (1988) showed that methylated, long-chain cellulose strongly inhibited cellulose degradation by several species of cellulolytic bacteria of ruminal origin. Also, other studies reported that methylcellulose inhibited attachment of cellulolytic rumen bacteria to cellulose fiber, which subsequently influenced fiber degradation (Rasmussen et al., 1989; Bhat et al. 1990). However, there are not any data so far that shows the relationship between bacterial attachment and fiber digestion by inhibition of methylcellulose. Our study has clearly and directly showed that fiber degradation was inhibited by methylcellulose following the same trend as attachment inhibition in major cellulolytic bacteria. Both inhibition of bacterial attachments and prevention of fiber digestion occurred concurrently in the initial incubation time and continued during the incubation. Also, the difference in DM digestibility between control and MC addition became larger with increasing incubation time, same as bacterial attachment to rice straw. This finding showed that the cause of inhibition of attachment was due to loss of cellulolytic enzyme activity as well as prevention of bacterial proliferation by methylcellulose. The inhibition of cellulolytic enzyme activity and bacterial growth by MC was previously reported by Rasmussen et al. (1988) and White et al. (1988).
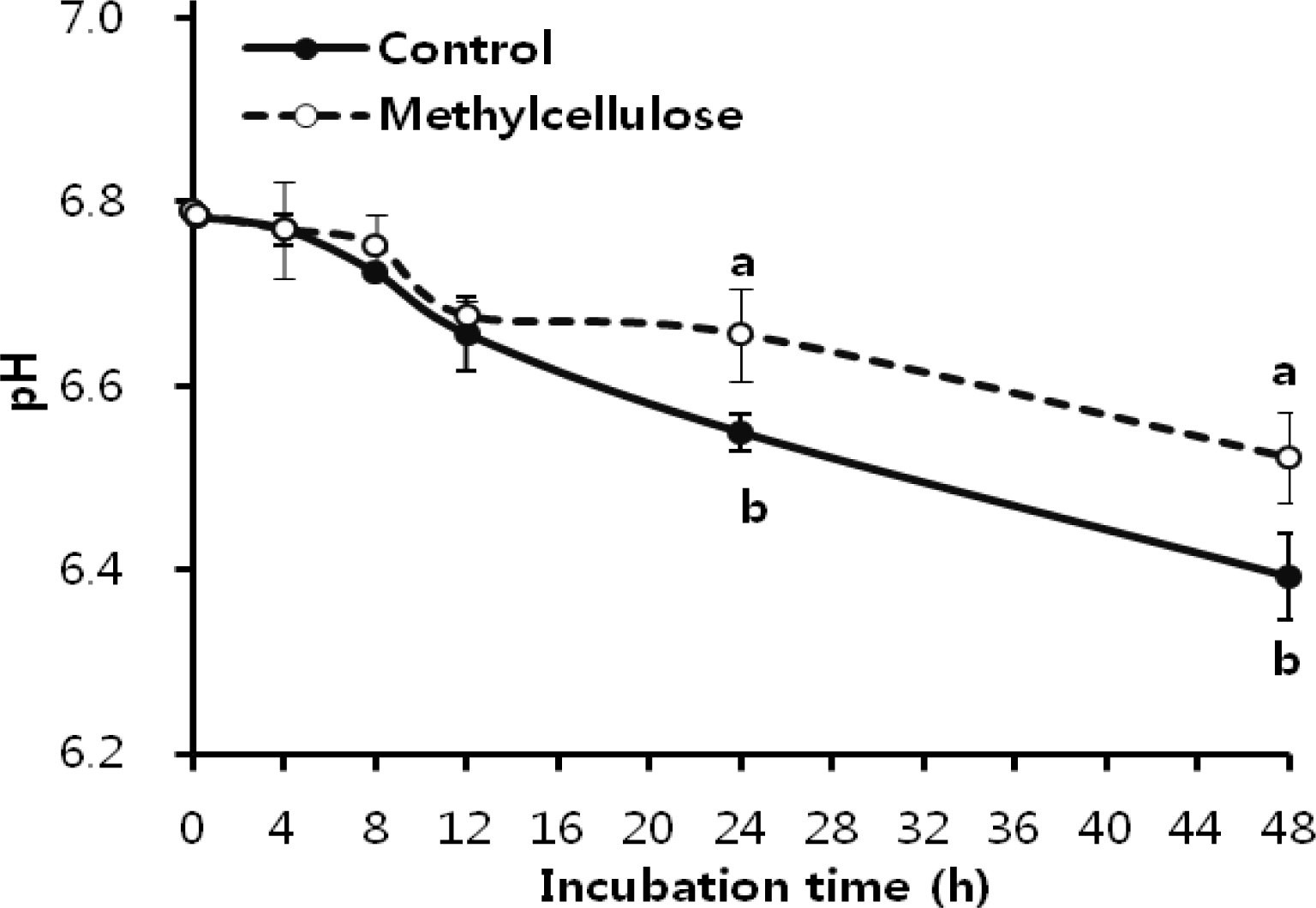
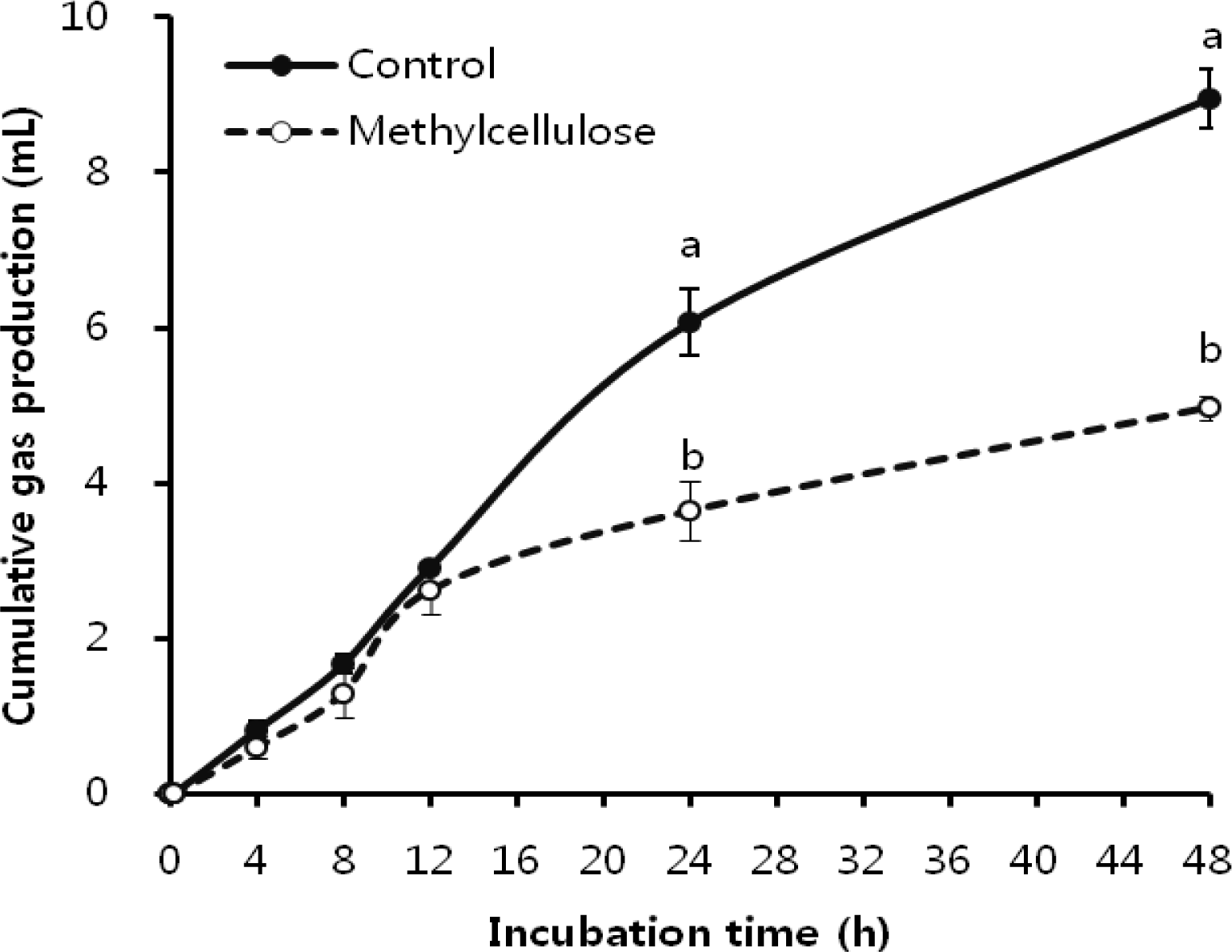
pH and cumulative total gas production during the incubation is shown in Figure 6 and 7. The pH showed similar decreasing trends until 12 h incubation. After 12 h incubation, pH showed a significant difference compared with control (p<0.05). Also, cumulative total gas production gradually increased with incubation time. Gas production showed similar increasing trends until 12 h incubation, and then there was a significant difference between methylcellulose treatment and control (p<0.05). Therefore this study suggests that methylcellulose inhibited bacterial attachments which subsequently decreased cellulose degradation; in addition it leads to abnormal fermentation in the rumen such as a decrease in gas production and pH decline.
ACKNOWLEDGMENTS
This research was supported by Bioindustry Technology Development Program, Ministry of Agriculture, Food & Rural Affairs, and Cooperative Research Program for Agriculture Science & Technology Development, Rural Development Administration, Republic of Korea.
Figure 1
The number of attached Fibrobacter succinogenes on rice straw as influenced by 0.1% methylcellulose. a,b Means with different letters differ significantly (p<0.05).
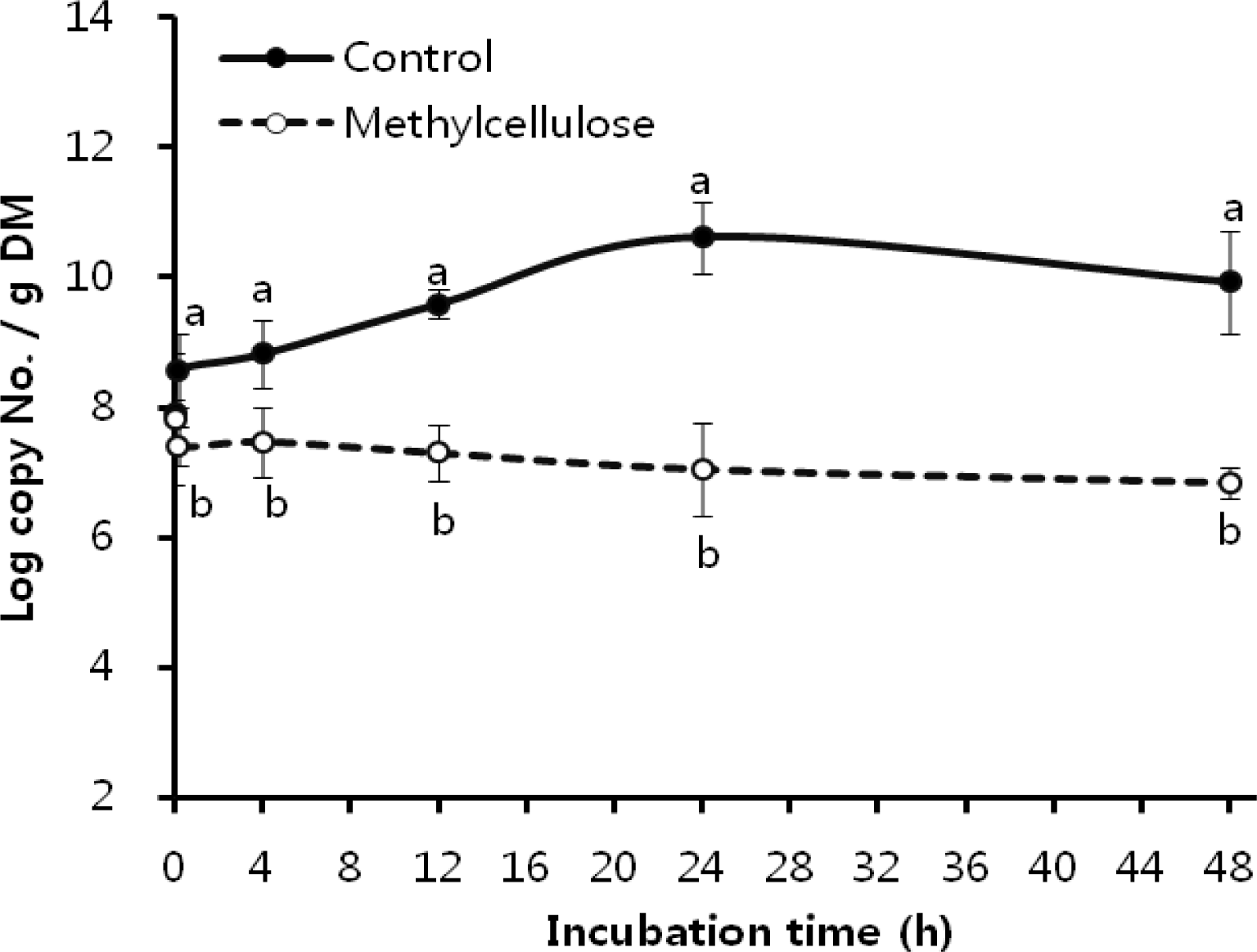
Figure 2
The number of attached Ruminococcus albus on rice straw as influenced by 0.1% methylcellulose. a,b Means with different letters differ significantly (p<0.05).

Figure 3
The number of attached Ruminococcus flavefaciens on rice straw as influenced by 0.1% methylcellulose. a,b Means with different letters differ significantly (p<0.05).
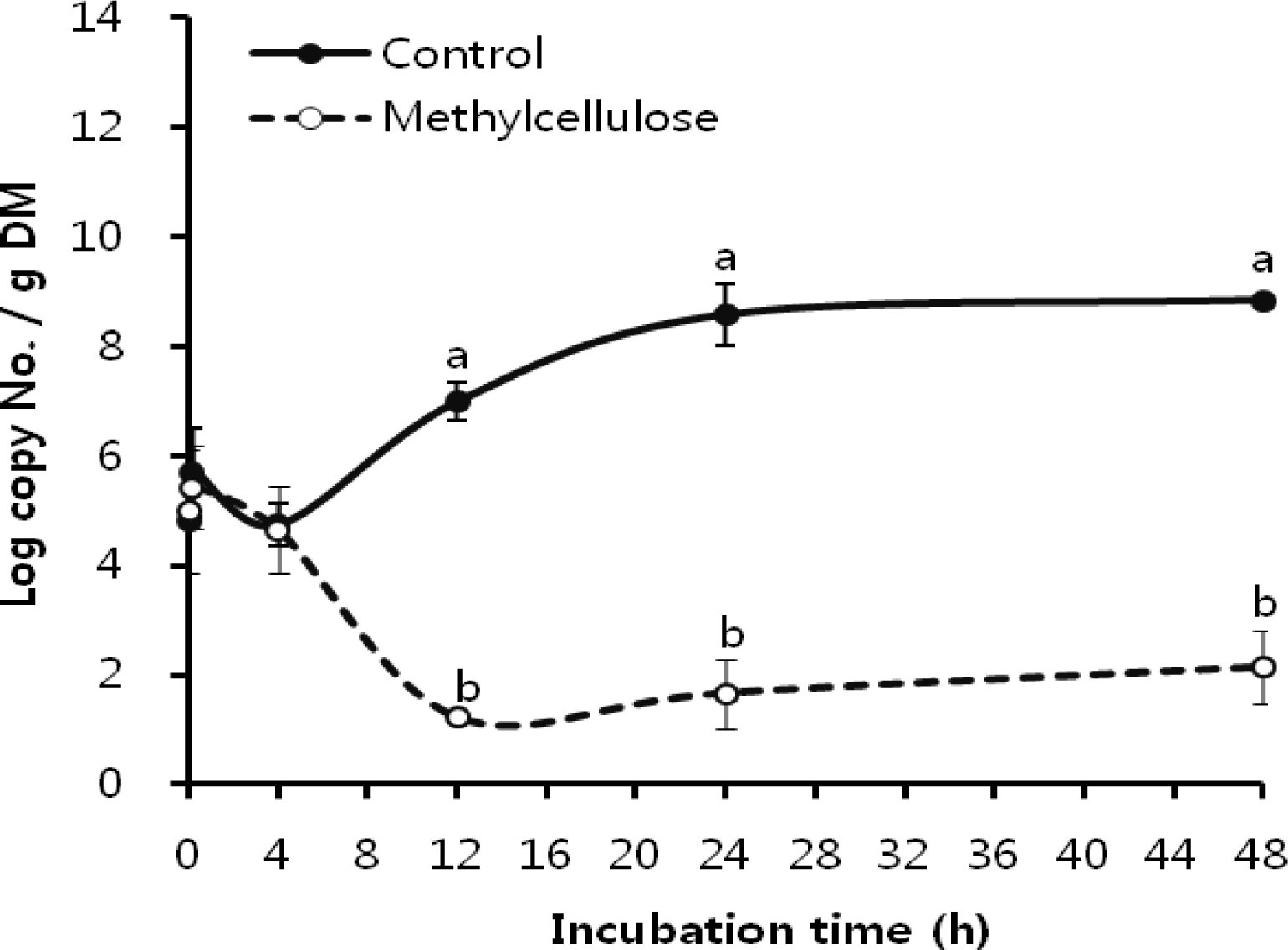
Figure 4
Fibrobacter succinogenes attachment on rice straw with methylcellulose in the early stage of incubation. a,b Means with different letters differ significantly (p<0.05).
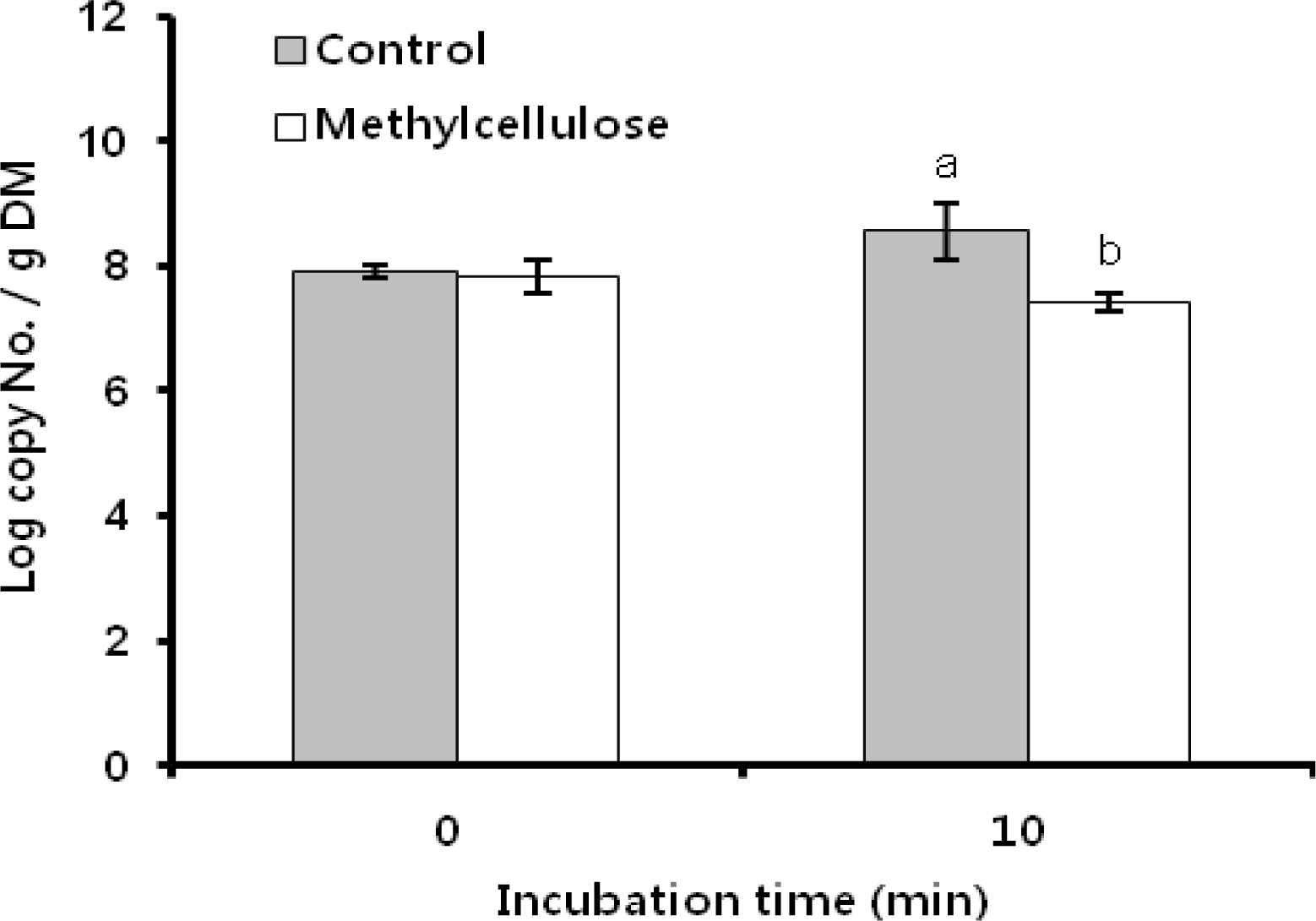
Figure 5
DM digestibility of rice straw as influenced by 0.1% methylcellulose. a,b Means with different letters differ significantly (p<0.05).
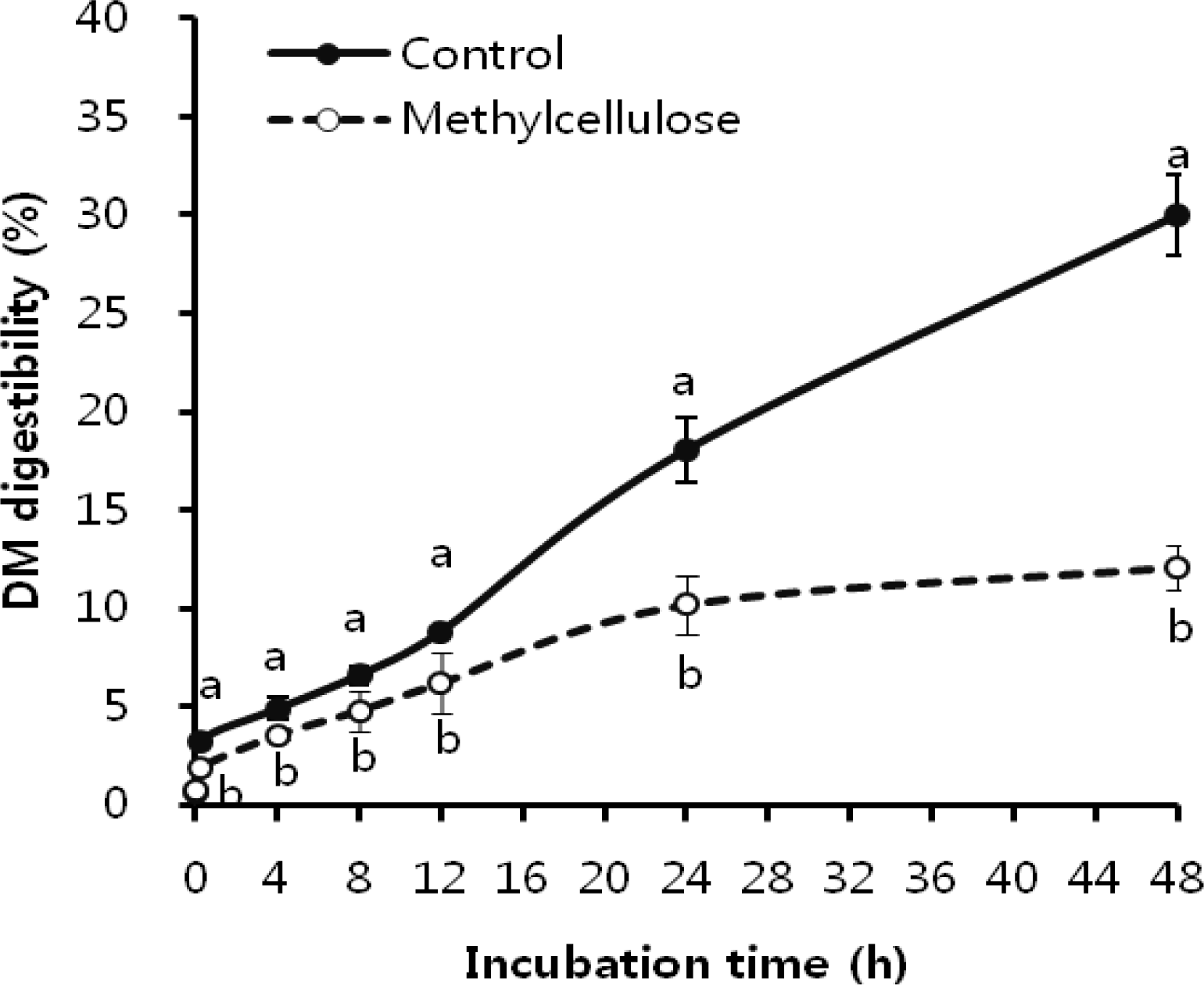
REFERENCES
Bhat S, Wallace RJ, Ørskov ER. 1990. Adhesion of cellulolytic ruminal bacteria to barley straw. Appl Environ Microbiol 56:2698–2703.



Cheng K-J, Stewart CS, Dinsdale D, Costerton JW. 1983. Electron microscopy of bacteria involved in the digestion of plant cell walls. Anim Feed Sci Technol 10:93–120.

Craig WM, Broderick GA, Ricker DB. 1987. Quantitation of microorganisms associated with the particulate phase of ruminal ingesta. J Nutr 117:56–62.


Czerkawski JW. 1986. Degradation of solid feeds in the rumen: spatial distribution of microbial activity and its consequences. Control of Digestion and Metabolism in Ruminants Proceedings of the Sixth International Symposium on Ruminant Physiology; Banff, Canada. p. 158–172. In : Milligan LP, Grovum WL, Dobson A, editorsEnglewood Cliffs, New Jersey: Prentice Hall.
Dinsdale D, Morris EJ, Bacon JSD. 1978. Electron microscopy of the microbial populations present and their modes of attach on various cellulosic substrates undergoing digestion in the sheep rumen. Appl Environ Microbiol 36:160–168.



Forsberg CW, Forano E, Chesson A. 2000. Microbial adherence to the plant cell wall and enzymatic hydrolysis. Ruminant Physiology: Digestion, Metabolism, Growth and Reproduction. p. 79–97. Cronje PB, editorWallingford: CABI Publishing.

Gong J, Forsberg CW. 1989. Factors affecting adhesion of Fibrobacter succinogenes S85 and adherence defective mutants to cellulose. Appl Environ Microbiol 55:3039–3044.



Hwang IH, Lee CH, Kim SW, Sung HG, Lee SY, Lee SS, Hong H, Kwan Y-C, Ha JK. 2008. Effects of mixtures of Tween80 and cellulolytic enzymes on nutrient digestion and cellulolytic bacterial adhesion. Asian-Aust J Anim Sci 21:1604–1609.

Koike S, Kobayshi Y. 2001. Development and use of competitive PCR assays for the rumen cellulolytic bacteria: Fibrobacter succinogenes, Ruminococcus albus and Ruminococcus flavefaciens. FEMS Microbiol Lett 204:361–366.


Kudo H, Cheng K-J, Costerton JW. 1987. Electron microscopic study of the methylcellulose-mediated detachment of cellulolytic rumen bacteria from cellulose fibers. Can J Microbiol 33:267–272.


Lee SS, Ha JK, Cheng K-J. 2000. Relative contributions of bacteria, protozoa, and fungi to in vitro degradation of orchard grass cell walls and their interactions. Appl Environ Microbiol 66:3807–3813.



McAllister TA, Bae HD, Jones GA, Cheng K-J. 1994. Microbial attachment and feed digestion in the rumen. J Anim Sci 72:3004–3018.


McDougall EI. 1948. Studies on ruminant saliva. 1. The composition and output of sheep’s saliva. Biochem J 43:99–109.


Michalet-Doreau B, Fernandez I, Peyron C, Millet L, Fonty G. 2001. Fibrolytic activities and cellulolytic bacterial community structure in the solid and liquid phases of rumen contents. Reprod Nutr Dev 41:187–194.


Minato H, Endo A, Ootome Y, Uemura T. 1966. Ecological treatise on the rumen fermentation. II. The amylolytic and cellulolytic activities of fractionated bacterial portions attached to the rumen solids. J Gen Appl Microbiol 12:53–61.

Minato H, Suto T. 1978. Technique for fractionation of bacteria in rumen microbial ecosystem. II. Attachment of bacteria isolated from bovine rumen to cellulose powder in vitro and elution of bacteria attached therefrom. J Gen Appl Microbiol 24:1–16.

Minato H, Suto T. 1979. Technique for fractionation of bacteria in the rumen microbial ecosystem. III. Attachment of bacteria isolated from bovine rumen to starch granules in vitro and elution of bacteria attached therefrom. J Gen Appl Microbiol 25:71–78.

Minato H, Mitsumori M, Cheng K-J. 1993. Attachment of microorganisms to solid substrates in the rumen. p. 139–145. Proc. MIE Bioforum on Genetics, Biochemistry and Ecology of Lignocellulose Degradation. Institut Pasteur; Paris, France:
Miron J, Forsberg CW. 1998. Features of Fibrobacter intestinalis DR7 mutant which is impaired with its ability to adhere to cellulose. Anaerobe 4:35–43.


Miron J, Forsberg CI. 1999. Characterization of cellulose binding proteins which are involved in adhesion mechanism of Fibrobacter intestinalis DR7. Appl Microbiol Biotechnol 51:491–497.


Miron J, Ben-Ghedalia D, Morrison M. 2001. Invited Review: Adhesion mechanisms of rumen cellulolytic bacteria. J Dairy Sci 84:1294–1309.


Morris EJ, Cole OJ. 1987. Relationships between cellulolytic activity and adhesion to cellulose in Ruminococus albus. J Gen Microbiol 133:1023–1032.
Pell AN, Schofield P. 1993. Microbial adhesion and degradation of plant cell walls. p. 397–423. Forage Cell Wall Structure and Digestibility. Hatfield RD, Jung HG, Ralph J, Buxton DR, Mertens DR, Weimer PJ, editorsASA-CSSA-SSSA; Madison, WI:

Purdy KJ, Embley TM, Takii S, Nedwell DB. 1996. Rapid extraction of DNA and rRNA from sediments by a novel hydroxyapatite spin-colum method. Appl Environ Microbial 62:3905–3970.

Rasmussen MA, White BA, Hespell RB. 1989. Improved assay for quantitating adherence of ruminal bacteria to cellulose. Appl Environ Microbiol 55:2089–2091.



Rasmussen MA, Hespell RB, White BA, Bothast RJ. 1988. Inhibitory effects of methylcellulose on cellulose degradation by Ruminococcus flavefaciens. Appl Environ Microbiol 54:890–897.



Reddy SKK, Morrison M. 1998. Biochemical and molecular characterization of adherence-defective mutants of Ruminococcus albus strain 8. p. 132Proc. MIE Bioforum on Cellulose Degradation. Institut Pasteur; Paris, France:
Statistical Analysis System Institute. 1996. SAS/STATTM User’s Guide: Statistics, Version 7. 5th EditionCary, NC:
Stewart CS, Duncan SH, Flint HJ. 1990. The properties of forms of Ruminococcus flavefaciens which differ in their ability to degrade cotton cellulose. FEMS Microbiol Lett 72:47–50.

Sung HG, Kobayashi Y, Chang J, Ha A, Hwang IH, Ha JK. 2007. Low ruminal pH reduces dietary fiber digestion via reduced microbial attachment. Asian-Aust J Anim Sci 20:200–207.

Trabalza-Marinucci M, Poncet C, Delval E, Fonty G. 2006. Evaluation of techniques to detach particle-associated microorganisms from rumen contents. Anim Feed Sci Technol 125:1–16.






 PDF Links
PDF Links PubReader
PubReader ePub Link
ePub Link Full text via DOI
Full text via DOI Full text via PMC
Full text via PMC Download Citation
Download Citation Print
Print





